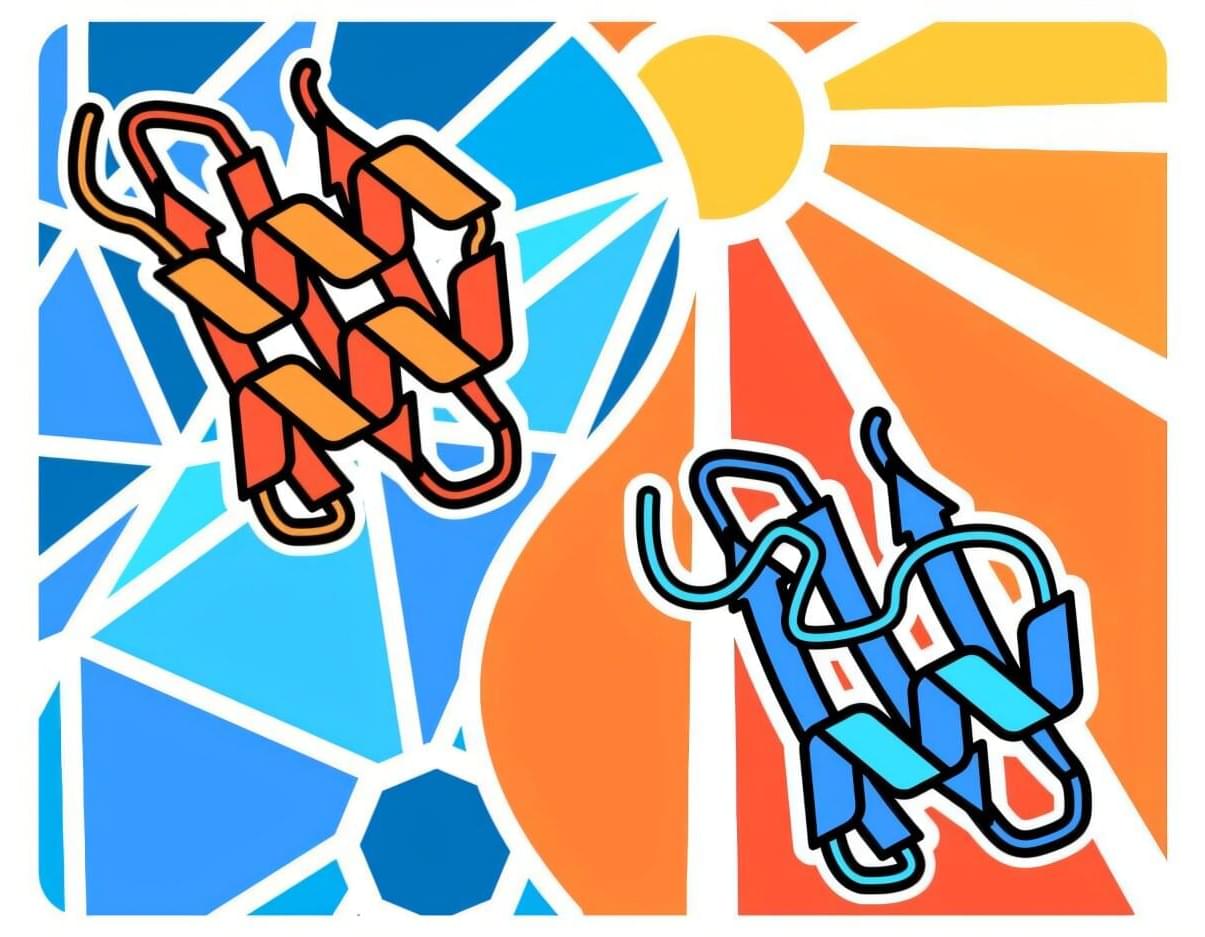
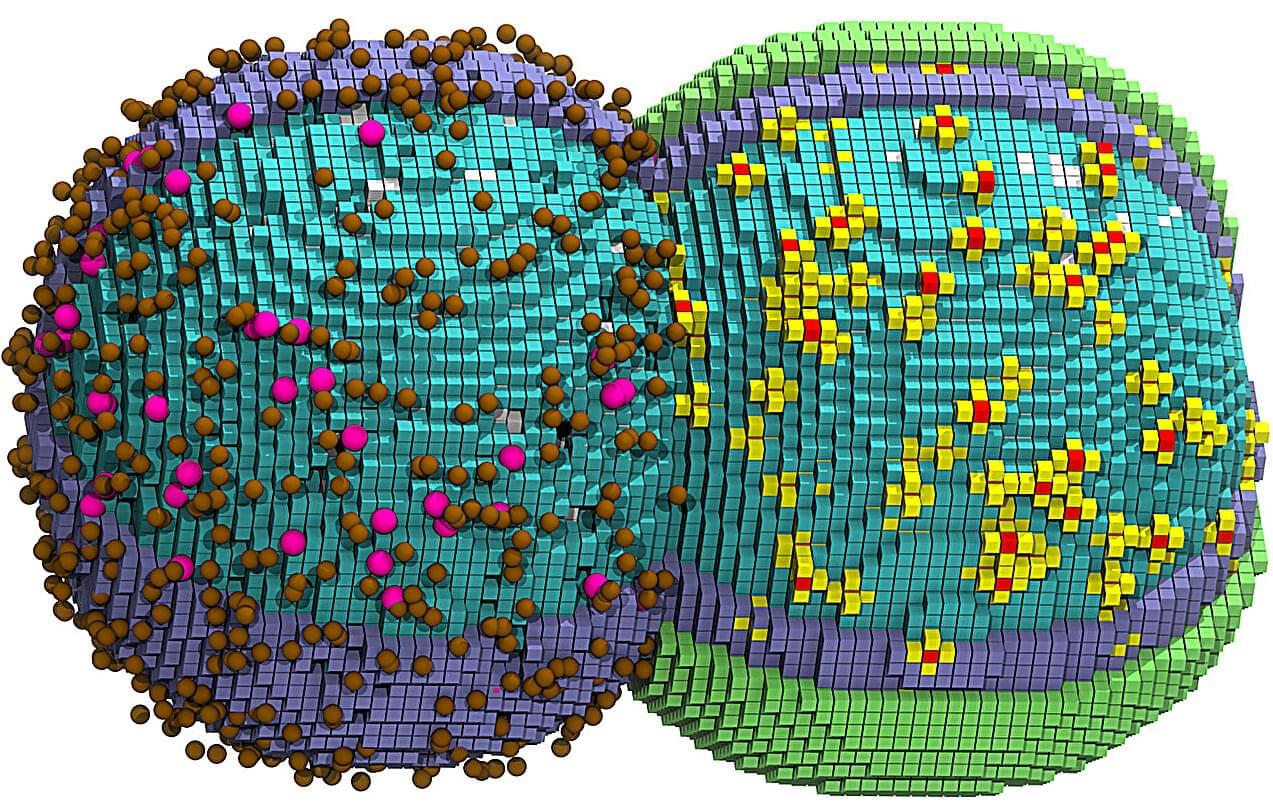
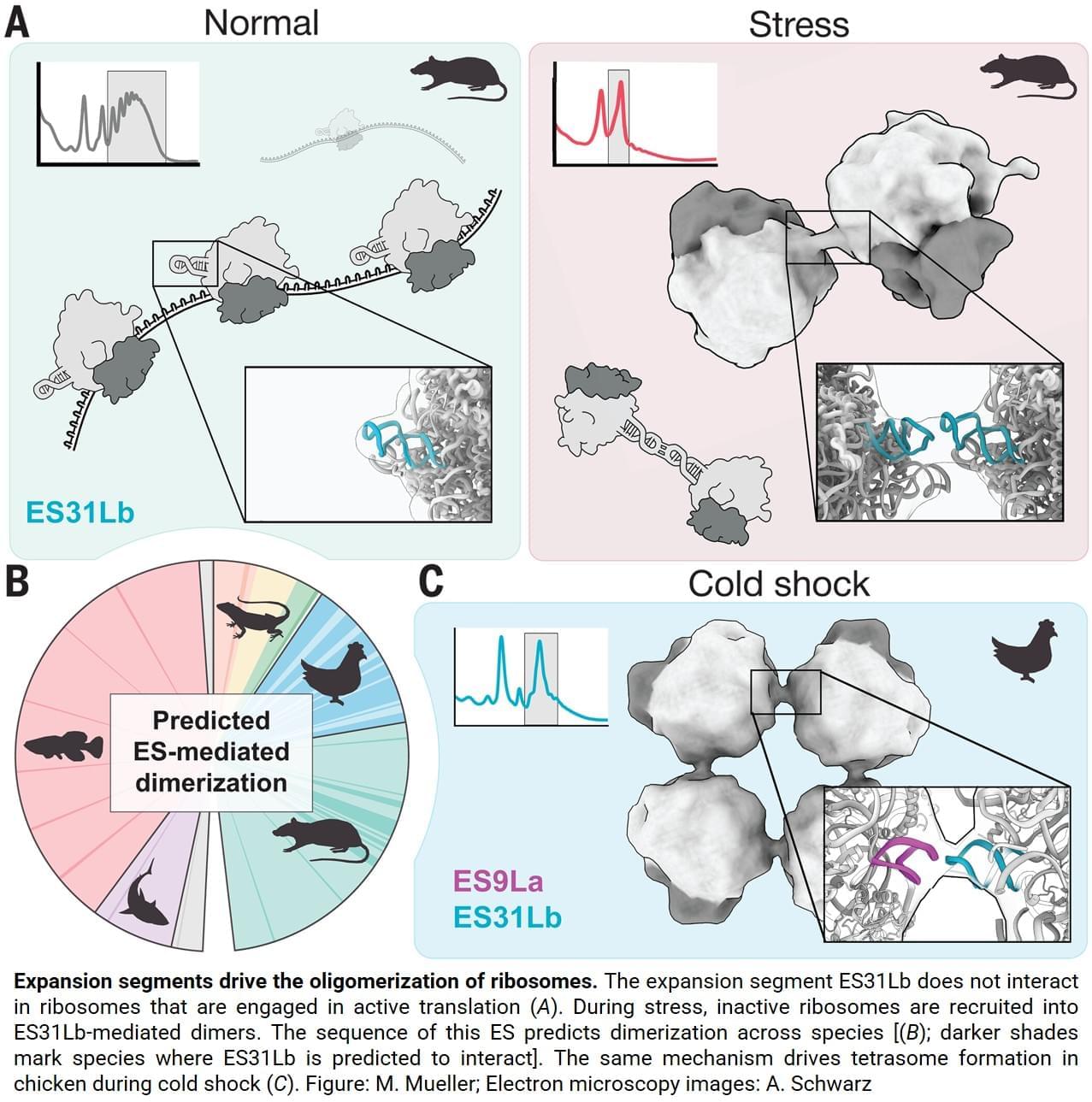
“Surprisingly, the two ribosomes are not held together by proteins, as is common in bacteria. Instead, the connection is made by a specific piece of ribosomal RNA called an expansion segment”, explains one of the lead authors.
Expansion segments are long, flexible RNA “tentacles” that protrude from ribosomes and have grown larger over the course of evolution. Although they are a prominent feature of animal ribosomes, their functions only just started to emerge. This study now shows that one particular expansion segment, called “31b”, is both necessary and sufficient to link ribosomes together during stress. At the molecular level, the expansion segment forms a precise RNA-RNA interaction — a so-called “kissing loop” — in which identical RNA loops bind each other through complementary sequences. Disrupting this interaction prevents disome formation, stunts cellular growth and makes cells more sensitive to stress. Science Mission sciencenewshighlights.
Ribosomes, the cell’s protein-making factories, consume large amounts of energy as they build the proteins that keep cells alive and functioning. When cells experience stress — such as lack of nutrients or sudden drops in temperature — they quickly switch into survival mode. New research now reveals an unexpected way cells manage this transition: By pairing up inactive ribosomes using a ribosomal RNA link. This RNA-based mechanism reveals a previously unknown role for ribosomal RNA in the cellular stress response.
Ribosomes are large molecular machines made of protein and RNA that build all proteins in the cell. Because protein production is extremely energy-intensive, cells rapidly reduce protein synthesis when stressed. It has long been known that bacterial cells pair their inactive ribosomes into so-called “hibernating disomes” however, such structures had not previously been identified in animal cells.
Using advanced imaging techniques, the team discovered that stressed animal cells — including neurons — assemble inactive ribosomes into tightly linked pairs, known as disomes. These ribosome pairs are not accidental collisions or artifacts, but a regulated and reversible response to stress. The new study was published in Science.