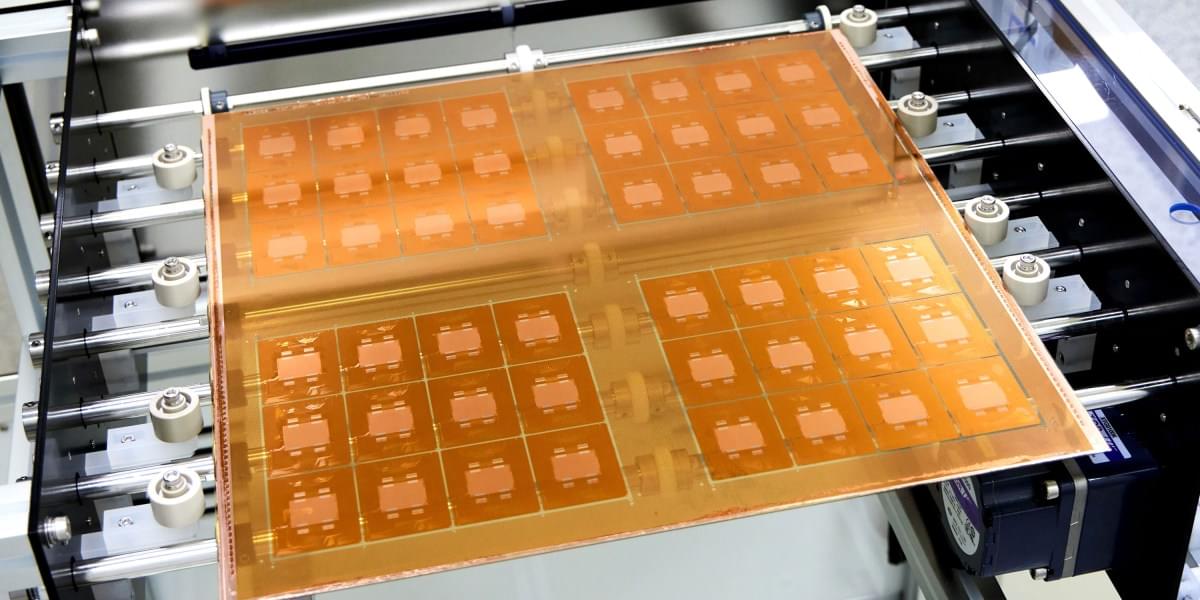
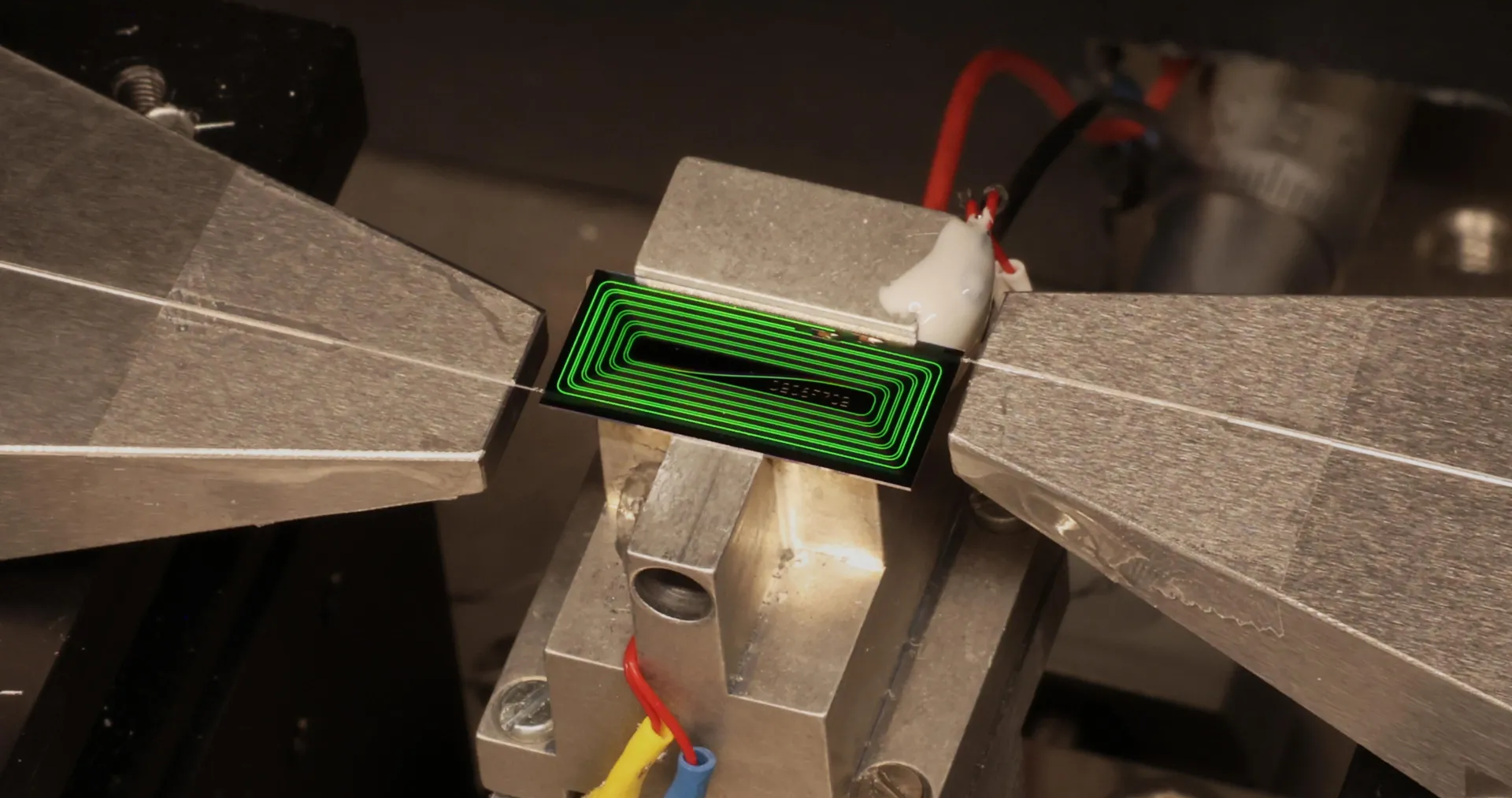
The idea is to use glass as the substrate, or layer, on which multiple silicon chips are connected. This form of “packaging” is an increasingly popular way to build computing hardware, because it lets engineers combine specialized chips designed for specific functions into a single system. But it presents challenges, including the fact that hardworking chips can run so hot they physically warp the substrate they’re built on. This can lead to misaligned components and may reduce how efficiently the chips can be cooled, leading to damage or premature failure.
“As AI workloads surge and package sizes expand, the industry is confronting very real mechanical constraints that impact the trajectory of high-performance computing,” says Deepak Kulkarni, a senior fellow at the chip design company Advanced Micro Devices (AMD). “One of the most fundamental is warpage.”
That’s where glass comes in. It can handle the added heat better than existing substrates, and it will let engineers keep shrinking chip packages—which will make them faster and more energy efficient. It “unlocks the ability to keep scaling package footprints without hitting a mechanical wall,” says Kulkarni.