KU Leuven, Belgium bioscience engineers have developed a roadmap, so to speak, for industrial cellulose gasoline.
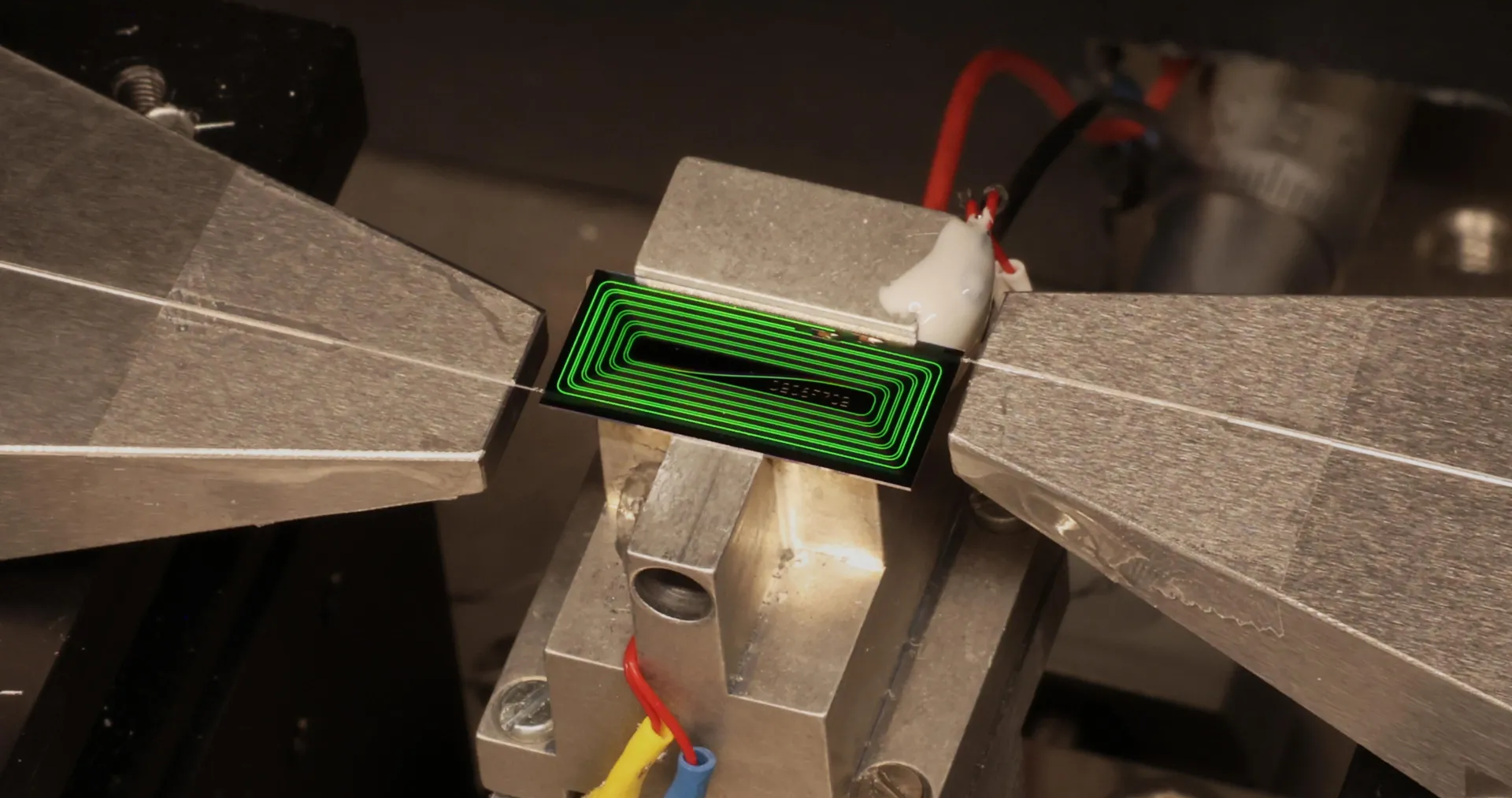
The bioscience engineers already knew how to make gasoline in the laboratory from plant waste such as sawdust. In 2014, at KU Leuven’s Centre for Surface Chemistry and Catalysis, the researchers succeeded in converting sawdust into building blocks for gasoline.
A chemical process made it possible to convert the cellulose – the main component of plant fibers – in the sawdust into hydrocarbon chains. These hydrocarbons can be used as an additive in gasoline. The resulting cellulose gasoline is a second-generation biofuel.