The hair cells lining the inner ear are among the most sophisticated structures in the human body: capable of detecting sounds as faint as a whisper, while helping to maintain our sense of balance. Through new models detailed in PRX Life, a team led by Roman Belousov at the European Molecular Biology Laboratory has revealed for the first time how oscillating bundles attached to these cells operate in different thermodynamic regimes—offering a new framework for understanding how our hearing works at a fundamental level.
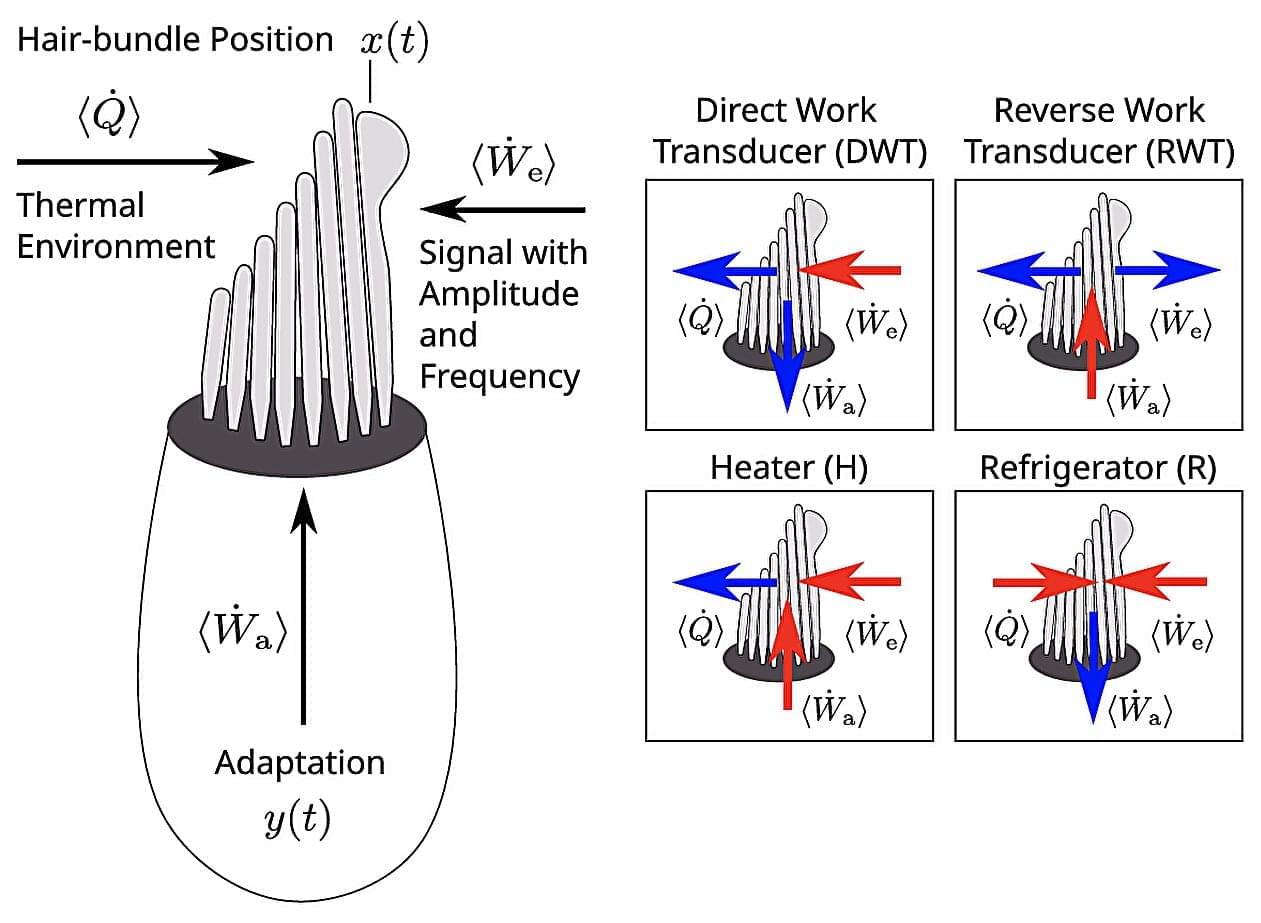
Within the inner ear, each hair cell hosts a hair “bundle”: a cluster of tiny, bristle-like projections that vibrate in response to incoming sound waves. The mechanical energy from these oscillations is then converted into electrical signals which travel to the brain. Rather than being passive receivers, these bundles actively oscillate —driven by molecular motors within the cell that allow them to amplify faint signals and tune in to specific frequencies.
But despite decades of study, researchers are still unclear on the connection between this active oscillation and the hair bundle’s response to external sound. Existing models tended to treat bundles as if they were moving spontaneously, without accounting for what happens when they actually interact with sound.