A personalized vaccine to treat glioblastoma, a fast-growing and incurable brain cancer that affects four in 100,000 people in the U.S., is safe and elicits robust and broad immune responses that appears to increase recurrence-free survival in a subset of patients after surgery, according to an early-stage clinical trial co-led by researchers at Washington University School of Medicine in St. Louis.
In patients with an especially aggressive form of glioblastoma, the vaccine caused no serious side effects and prolonged patients’ overall survival compared to historical outcomes after standard-of-care surgery and chemo-radiotherapy. One long-term survivor remains recurrence-free nearly five years later.
The results of the phase 1 trial, conducted at Siteman Cancer Center, based at Barnes-Jewish Hospital and WashU Medicine, were published May 12 in Nature Cancer. The study was led jointly by Mass General Brigham and Geneos Therapeutics, a Philadelphia-based biotechnology company.
“We are extremely encouraged by these results,” said Tanner M. Johanns, MD, PhD, lead author of the study and an assistant professor in the Division of Oncology in the John T. Milliken Department of Medicine at WashU Medicine. “This kind of vaccine is a first for glioblastoma, and it is exciting to think how we can leverage this individualized therapeutic DNA cancer vaccine platform to make a positive impact on the lives of patients who are fighting this disease. Additionally, combination therapies leveraging this personalized platform are currently being investigated at WashU to test if outcomes may be improved further.”
Abstract: Nature Cancer
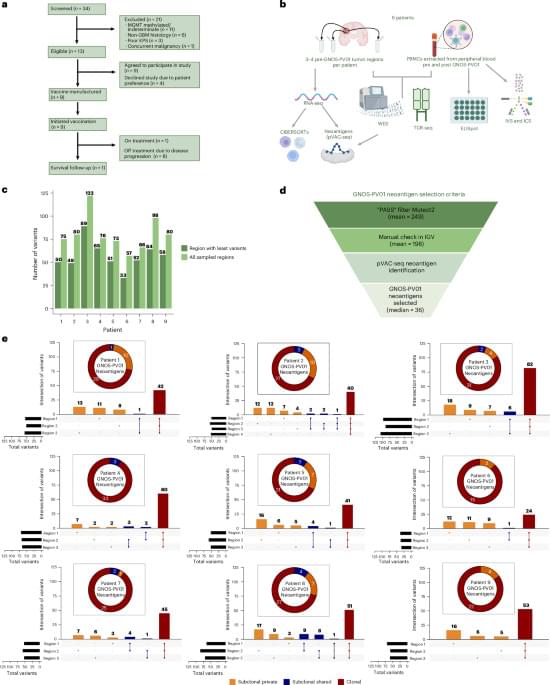
Johanns and colleagues report the results (including safety, efficacy and immunogenicity) of a phase 1 clinical trial of a DNA-based personalized therapeutic cancer vaccine administered following surgical resection and radiation in patients with MGMT unmethylated glioblastoma.