The crucial drugs can have unintended consequences. Innovative therapies could shield the microbiome from their effects.
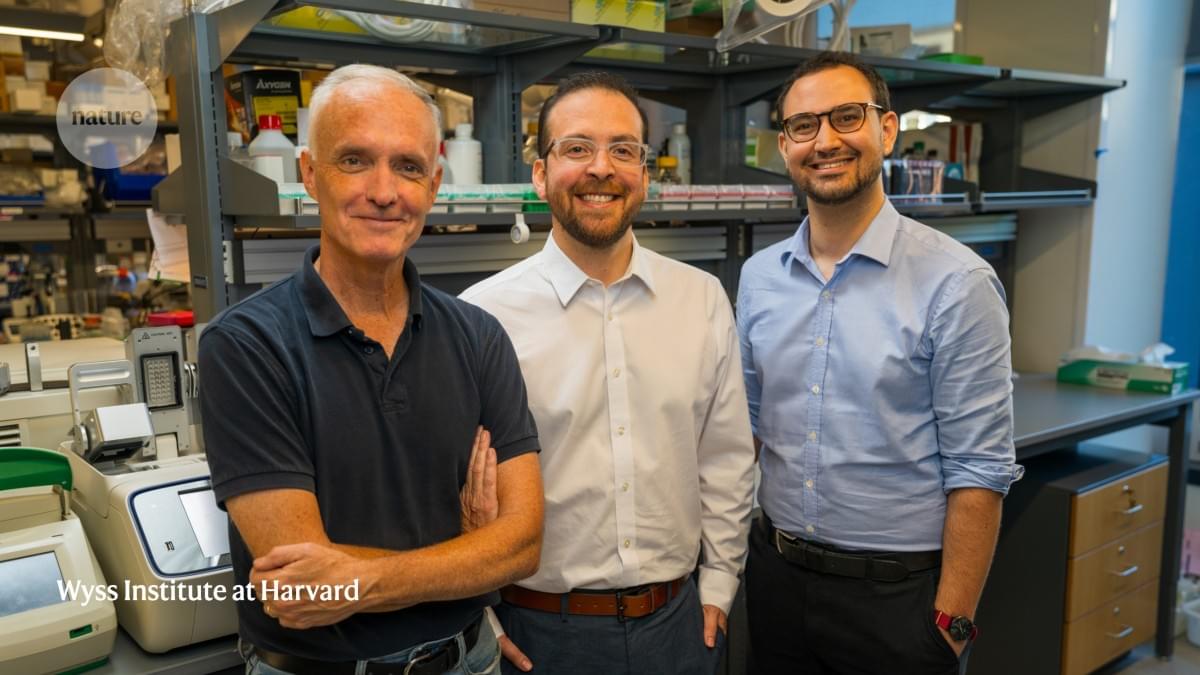
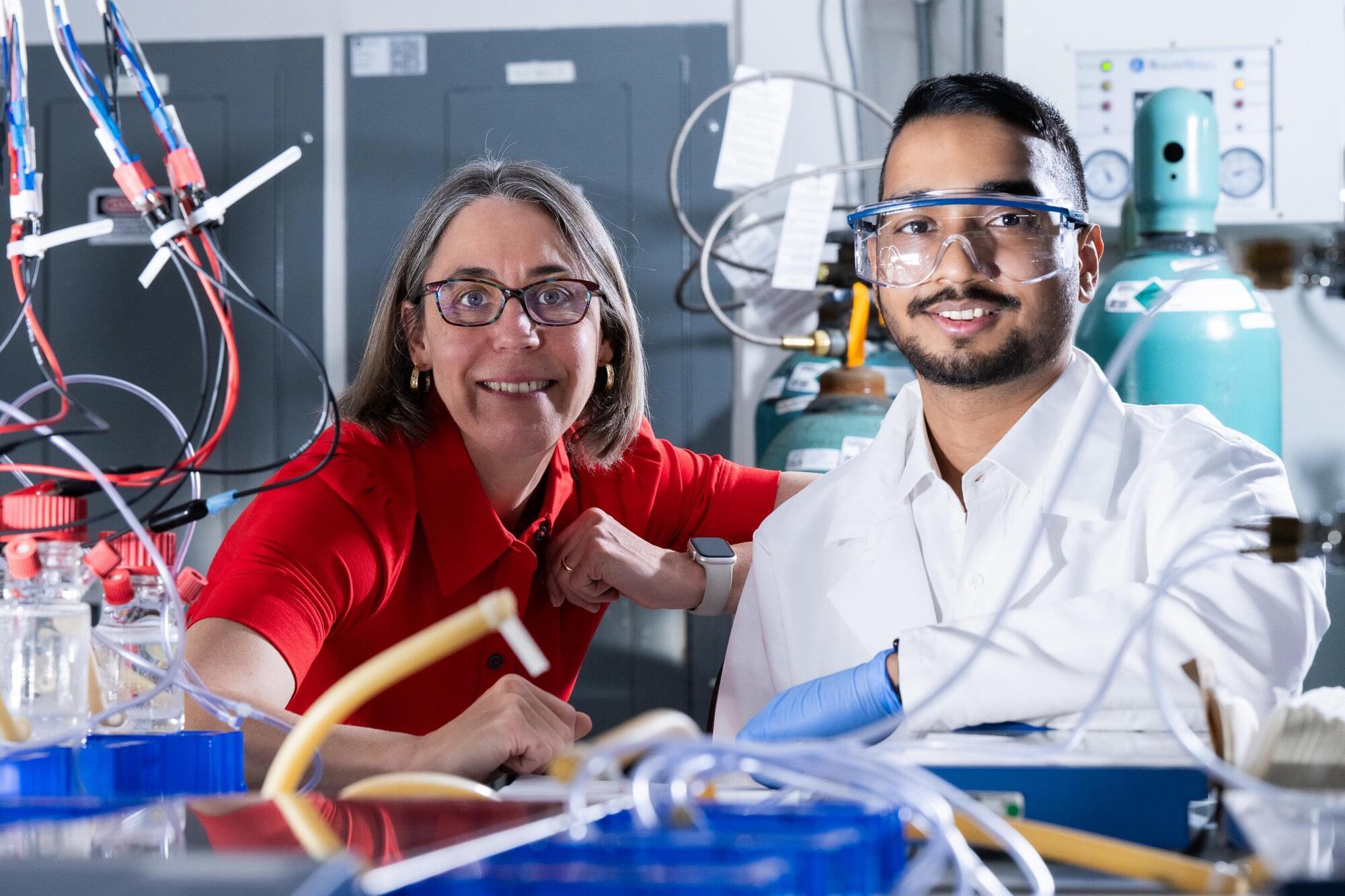



A team led by Rice University bioscientist Caroline Ajo-Franklin has discovered how certain bacteria breathe by generating electricity, using a natural process that pushes electrons into their surroundings instead of breathing on oxygen.
The findings, published in Cell, could enable new developments in clean energy and industrial biotechnology.
By identifying how these bacteria expel electrons externally, the researchers offer a glimpse into a previously hidden strategy of bacterial life. This work, which merges biology with electrochemistry, lays the groundwork for future technologies that harness the unique capabilities of these microscopic organisms.
Many diseases are caused by a missing or defective copy of a single gene. For decades, scientists have been working on gene therapy treatments that could cure such diseases by delivering a new copy of the missing genes to the affected cells.
Despite those efforts, very few gene therapy treatments have been approved by the FDA. One of the challenges to developing these treatments has been achieving control over how much the new gene is expressed in cells — too little and it won’t succeed, too much and it could cause serious side effects.
To help achieve more precise control of gene therapy, MIT engineers have tuned and applied a control circuit that can keep expression levels within a target range. In human cells, they showed that they could use this method to deliver genes that could help treat diseases including fragile X syndrome, a disorder that leads to intellectual disability and other developmental problems.


Over the past several decades, human lifespan has steadily increased. However, this progress has also led to a growing proportion of the population suffering from age-related diseases such as cancer, neurodegenerative disorders, and diabetes. Extending both lifespan and healthspan, the period of life spent in good health, requires a deeper understanding of the biological mechanisms that promote healthy aging.
In the natural world, mammalian lifespans vary enormously, ranging from just 1 to 2 years in some rodents to more than a century in species.
A species is a group of living organisms that share a set of common characteristics and are able to breed and produce fertile offspring. The concept of a species is important in biology as it is used to classify and organize the diversity of life. There are different ways to define a species, but the most widely accepted one is the biological species concept, which defines a species as a group of organisms that can interbreed and produce viable offspring in nature. This definition is widely used in evolutionary biology and ecology to identify and classify living organisms.
In a study published Wednesday in the Proceedings of the National Academy of Sciences, University of Oklahoma researchers detail their discoveries about why the brain tumor glioblastoma is so aggressive. Their findings center on ZIP4, a protein that transports zinc throughout the body and sets off a cascade of events that drive tumor growth.
About half of all malignant brain tumors are glioblastomas, the deadliest form of brain cancer with a median survival rate of 14 months.
“Surgery for glioblastoma is very challenging, and patients almost always experience a relapse,” said the study’s senior author, Min Li, Ph.D., a professor of medicine, surgery and cell biology at the University of Oklahoma College of Medicine. “By better understanding why these brain tumors are so aggressive, we hope to open up paths for new treatments.”

The immune system is comprised of two separate active arms of immunity to provide robust protection against disease. The two separate systems of immunity include the innate and adaptive immune responses. The innate immune system is the first on the scene when a pathogen enters the body. Different cells of this response include eosinophils, basophils, neutrophils, natural killer cells, macrophages, dendritic cells, and others. Once a pathogen is detected the innate immune system generates a generalized response to target a wide variety of diseases. The innate immune cells then relay detection of a foreign pathogen to the second level of immunity, the adaptive immune response. The second wave that fights off disease is more specific and includes T and B cells. The adaptive immune response is widely accepted as the stronger barrier of immunity because of its specificity and ability to mount a larger response. However, each work in concert with the other to elicit an optimal immune response and keep the body healthy.
Many different cells are involved in active immunity that help protect against infection. As a result, immunologists work to understand each cell’s impact on the immune system and how they function in the context of various pathologies. To activate the immune system during an infection or disease onset, immunotherapy is employed, which targets one or several immune cell types to redirect them to the infection. In particular, natural killer cells, are emerging as an optimal cell target for immunotherapy in ovarian cancer.
Natural killer cells are a type of white blood cell responsible for eliminating virus-infected cells and tumor cells. They secrete various molecules and proteins that signal a strong immune response. Additionally, they have been reported to develop immune memory, in which they can recognize previously encountered pathogen. These natural killer cells that develop immune memory are referred to as adaptive natural killer (aNK) cells. Scientists are working to understand more about these cell types and how they can be targeted for cancer immunotherapy.


A Kobe University team was able to edit the DNA of Lactobacillus strains directly without a template from other organisms. This technique is indistinguishable from natural variation and enabled the researchers to create a strain that doesn’t produce diabetes-aggravating chemicals.
Humans have improved the microorganisms we rely on for millennia, selecting variants that are better able to produce wine, yogurt, natto and many other products. More recently, direct genetic modification has emerged as a tool to exert more precise and efficient control over the improvement, but also has drawn much public criticism for often using DNA from unrelated organisms in these modifications. Kobe University bioengineer NISHIDA Keiji says, “As a consequence, using such transgenic techniques is not favorable for food products due to legislations being restrictive and social acceptance being low.”
Nishida and his team have developed a technique that gives even more precise control over the genetic content of a microorganism that does not rely on template DNA from other organisms. He says: “We have invented a DNA base editing technology named ‘Target-AID,’ which is superior to conventional techniques such as ‘CRISPR-Cas9’ in several aspects. For example, CRISPR-Cas9 induces DNA breaks and often causes cell death, while our Target-AID inserts precise point mutations without such breaks.”

Researchers at Amsterdam UMC have developed a new diagnostic test that can quickly and accurately diagnose bacterial meningitis. The test measures the CRP protein in cerebrospinal fluid, a protein that is already often tested in blood to detect bacterial infections. Currently, it often takes a long time before meningitis is diagnosed, which delays the start of adequate treatment.
The study is published in The Lancet Regional Health—Europe.
Bacterial meningitis is a life-threatening condition in which one in six patients die and half of the survivors have residual symptoms. Thus, prompt diagnosis and treatment are crucial.